Stay tuned...
For more detailed information, please refer to the documentation.
To address the rapid growth of high-dimensional data in biology and other fields, we present MILK, a scalable approach for capturing highly granular relationships between objects (e.g., single cells). By recursively stratifying the population into groups of highly similar objects based on pairwise distance, MILK builds a hierarchical organization of objects that preserves both global structure and rare local subpopulations. This provides a framework for multi-resolution analysis of ultra-large datasets in a tractable and interpretable manner.
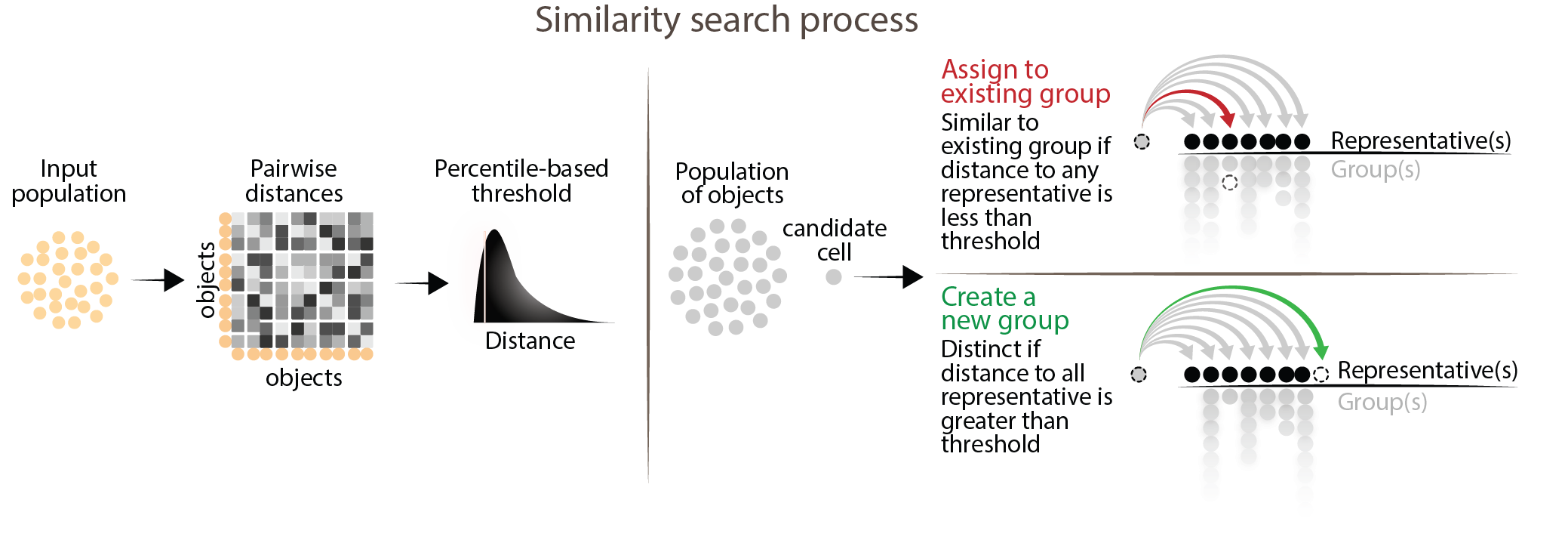
The main process of MILK involves grouping highly similar objects using a data-driven similarity threshold defined by a lower-tail percentile (e.g., 0.1%) of distances. In a single-pass through the dataset, each candidate object is compared to the representatives of existing groups. If the candidate has a distance below the threshold to any group, it is mapped to the group it is most similar to; otherwise, it is considered distinct, in which case it forms a new group and becomes its representative. The first candidate object forms a new group by default.
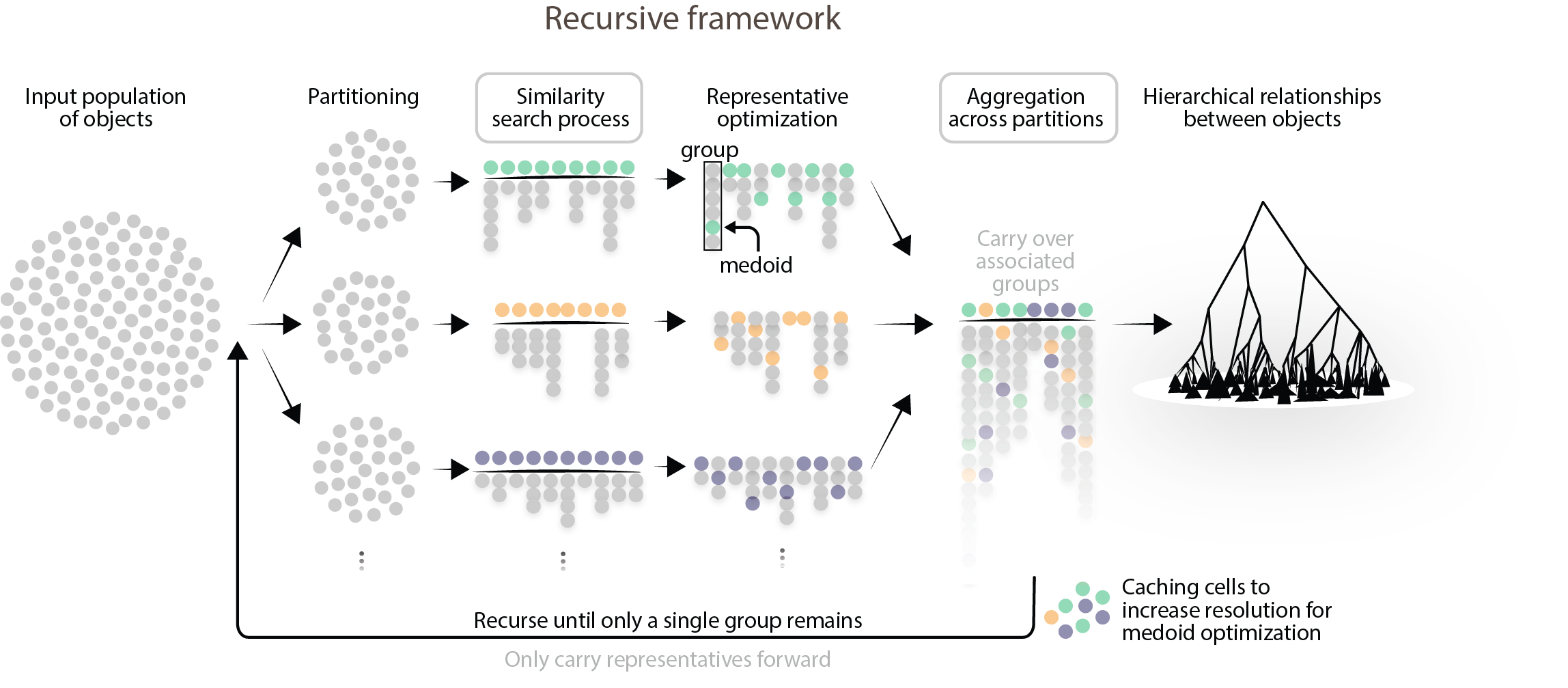
To handle datasets that exceed available memory, MILK employs a recursive, out-of-core strategy that involves partitioning the population into tractable subsets and processing them with a shared global similarity threshold. After identifying groups within each subpopulation, representatives are optimized to be the medoid of each group. All groups are then aggregated into a global list and, if tractable, an additional grouping process is performed on the representatives to resolve potential overlap across partitions.
By carrying forward only representatives, this pipeline can be applied recursively to progressively merge groups according to the percentile-based similarity threshold. If allowed to run to completion, a global hierarchical structure is produced over the complete population.
Users can run MILK without installing Julia on your system.
Important
Pre-release note: currently, the x86 (64-bit) build is provided1
Download the latest milk build from the Latest Releases page.
Extract binary:
tar -xzf milk-linux-x86_64.tar.gz
Export to PATH (replace /path/to with the actual download location):
export PATH="$PATH:/path/to/milk/bin"
Install Julia from the official website.
Note that MILK was developed and tested using Julia version 1.11.3.
Install MILK using Julia's package manager:
In REPL:
using Pkg
Pkg.add(url="https://github.com/yachielab/milk.git")
Or, from the shell:
julia -e 'using Pkg; Pkg.add(url="https://github.com/yachielab/milk.git")'
Optionally, users can export the source directory to PATH:
echo 'export PATH="$PATH:./milk"
Confirm successful installation with the following call:
milk -h
MILK can be applied to any high-dimensional dataset. It expects an uncompressed .CSV file format as input, with the first column corresponding to object IDs.
milk -i input.csv
The full set of arguments can be seen below:
usage: milk -i INPUT-PATH [-n SAMPLE-SIZE] [-c CACHE-SIZE-LIMIT]
[-p PARTITION-SIZE] [-M MERGE-THRESHOLD] [-m METRIC]
[-l LABEL] [-t PERCENTILE] [-T THREADS] [-r SEED]
[--verbose] [--force-overwrite] [--skip-reconstruction]
[-o OUTPUT-DIR] [--hpc-mode] [-b BATCH-SIZE]
[--job-scheduler JOB-SCHEDULER]
[--job-account JOB-ACCOUNT] [--job-name JOB-NAME]
[--job-memory JOB-MEMORY] [--job-time JOB-TIME]
[--environment-path ENVIRONMENT-PATH]
[--group-stratification-mode]
[--stratification-input-dir STRATIFICATION-INPUT-DIR]
[--stratification-threshold STRATIFICATION-THRESHOLD]
[--stratification-percentile STRATIFICATION-PERCENTILE]
[--stratification-metric STRATIFICATION-METRIC]
[--stratification-cache-path STRATIFICATION-CACHE-PATH]
[--stratification-previous-groups-path STRATIFICATION-PREVIOUS-GROUPS-PATH]
[--stratification-output-dir STRATIFICATION-OUTPUT-DIR]
[-h]
A command-line tool to capture hierarchical relationships at scale.
optional arguments:
-h, --help show this help message and exit
Main arguments:
-i, --input-path INPUT-PATH
Uncompressed CSV file. First column
corresponds to object IDs (e.g., cell
barcode). No header
-n, --sample-size SAMPLE-SIZE
Sample size threshold for the number of groups
required to stop the recursive grouping
process. Recursion stops when the number of
(representative) objects is less than or equal
to this value. (type: Int64, default: 1)
-c, --cache-size-limit CACHE-SIZE-LIMIT
How many (representative) objects to cache for
medoid optimization in subsequent recursions.
(type: Int64, default: 50000)
-p, --partition-size PARTITION-SIZE
How many objects per partitioned file. (type:
Int64, default: 10000)
-M, --merge-threshold MERGE-THRESHOLD
Threshold of when to carry out an additional
merging stratification process (if exceeded,
basic concatenation to join partitioned
groups) (type: Int64, default: 100000)
-m, --metric METRIC Distance metric for pairwise comparisons.
Permissible values:
{'cosine','euclidean','manhattan','hamming','jaccard','correlation'})
(default: "euclidean")
-l, --label LABEL Specify string label for file names.
-t, --percentile PERCENTILE
Threshold for stratification process to group
cells. (type: Float64, default: 1.0)
-T, --threads THREADS
Number of distributed processes for computing
pairwise comparisons. (TODO change to n_cpus)
(type: Int64, default: 1)
-r, --seed SEED Random seed. (type: Int64, default: 21)
--verbose Amount of information written to standard
out/err.
--force-overwrite Overwrite output directory if it already
exists.
--skip-reconstruction
Only perform hierarchical
grouping/downsampling phase and NOT the
additional step of reconstructing vertices and
edges table for graph structure.
-o, --output-dir OUTPUT-DIR
Directory to write intermediate and output
files. (default: "./milk.out")
[High-Performance Computing mode] Cluster computing for large-scale MILK runs:
--hpc-mode Activate HPC mode for MILK execution. If this
flag is not specified, all [HPC mode]
arguments will be ignored.
-b, --batch-size BATCH-SIZE
[HPC mode] The number of partitioned input
files to be assigned per job. (type: Int64,
default: 50)
--job-scheduler JOB-SCHEDULER
[HPC mode] Job scheduler to distribute
partitioned batches of stratification
processes. Supported schedulers:
{'slurm','sge'}
--job-account JOB-ACCOUNT
[HPC mode] Account name associated with
account (only for SLURM job scheduler).
--job-name JOB-NAME [HPC mode] Job names (default: "milk")
--job-memory JOB-MEMORY
[HPC mode] Memory of jobs (in gigabytes).
(type: Int64, default: 4)
--job-time JOB-TIME [HPC mode] Allotted time for jobs (only for
SLURM job scheduler). (default: "24:00:00")
--environment-path ENVIRONMENT-PATH
[HPC mode] Bash script to set up environment
in submitted jobs.