-
|
Hello, I created a wetland mask from a landsat image using qgis reclassify by table tool. This reclassification allowed me to create a binary raster of wetlands area. However if i run a pca or a kmeans using the mask i got data on the masked area. Exemple for pca : Steps :
Result : If i want to apply the mask, i have to use « Apply mask layer to raster layer », even if i already filled the mask in the « create unsupervised dataset » step. Is it normal ? Here is an extract of my data if you need them |
Beta Was this translation helpful? Give feedback.
Replies: 3 comments 1 reply
-
Beta Was this translation helpful? Give feedback.
-
|
Thank you, If i mask my original landsat_data.tif before drawing the samples then i obtain the following message when runnning "transform raster layer tool" : Traceback (most recent call last): Looks like a scikit issue... Workaround is to apply mask on output pca layers |
Beta Was this translation helpful? Give feedback.
-
|
Ok it works with the dev version |
Beta Was this translation helpful? Give feedback.



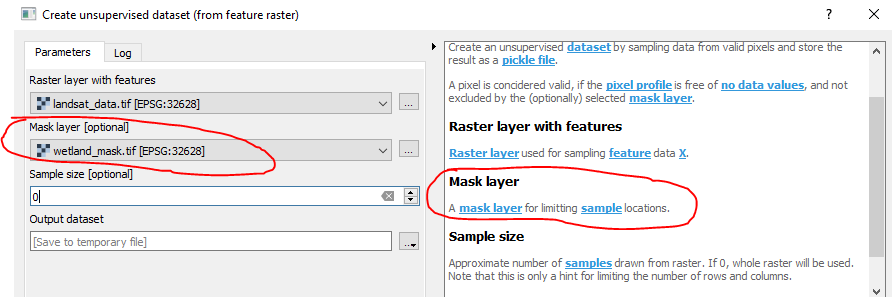
The mask that you specify here is only limitting the training samples for the PCA.
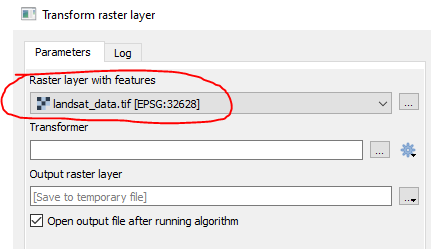
In the Transform step, only those areas are masked, which are already masked in the raster:
Note that instead of transforming the original raster, you could transform any other Landsat raster with the fitted PCA model. For those other rasters, the original mask would have no meaning.
Long story short. You could a) mask your original landsat_data.tif before drawing the samples, or b) mask the output PCA raster.